base excision repair
Base excision repair (BER) describes one form of excision repair in which damaged bases or incorrect bases are excised and replaced through specific enzymes that differ between species. However, the biochemical processes involved in BER are equivalent across species, so bacterial DNA repair functions can operate in eukaryotic cells, and vice versa. Damage is typically the result of deamination, alkylation, hydroxylation, or attack by an oxygen radical, while the incorrect base can be uracil substituted for thymine. Oxidative DNA lesions induced by oxygen free radicals such as superoxide and hydroxyl radicals appear to be repaired predominantly by base excision repair mechanisms. Further, BER is the major DNA repair system involved in removal of various oxidative DNA lesions induced by ionizing radiation - these include abasic sites and modified DNA base and sugar residues.
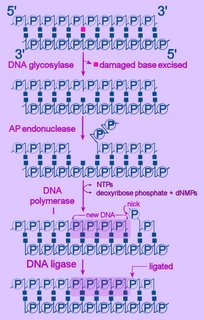 Left - diagram of base excision repair - click to enlarge image.
Left - diagram of base excision repair - click to enlarge image.
NTPs = ribonucleoside triphosphates
dNMP = deoxyribonucleoside monophosphate
First, the altered base is excised by a specific DNA glycosylase, which breaks the beta N-glycosidic bond and creates an AP, or abasic site. This site is identical to that generated by spontaneous depyrimidination or depurination. Six DNA glycosylases have been identified in humans – each excises an overlapping subset of either spontaneously formed (such as hypoxanthine), oxidized (such as 8-oxo-7,8-dihydroguanine), alkylated (such as 3-methyladenine), or mismatched (for example, T:G) bases.
Next, the terminal sugar-phosphate is removed by an AP endonuclease (Ape1), leaving a 3’-OH terminal and an abnormal 5'-abasic terminus. The resulting gap is refilled by the 5’-deoxyribose-phosphodiesterase action of a DNA polymerase I (DNA polymerase beta in mammals), then the strands are re-ligated by DNA Ligase I or a complex of XRCC1 and LigIII.
An alternative BER pathway corrects errors involving more than one nucleotide. The Fen1 protein excises the long-patch structure that is produced by DNA polymerase strand displacement. This "long-patch" repair process is divided into two subpathways: a PCNA-stimulated, Pol-beta-directed pathway and a PCNA-dependent, Pol-delta/epsilon -directed pathway.
link to table - human DNA repair genes : diagram>BER
Biochemistry Overview : Molecular Genetics Overview : SITE MAP : HOME

 Left - diagram of base excision repair - click to enlarge image.
Left - diagram of base excision repair - click to enlarge image.NTPs = ribonucleoside triphosphates
dNMP = deoxyribonucleoside monophosphate
First, the altered base is excised by a specific DNA glycosylase, which breaks the beta N-glycosidic bond and creates an AP, or abasic site. This site is identical to that generated by spontaneous depyrimidination or depurination. Six DNA glycosylases have been identified in humans – each excises an overlapping subset of either spontaneously formed (such as hypoxanthine), oxidized (such as 8-oxo-7,8-dihydroguanine), alkylated (such as 3-methyladenine), or mismatched (for example, T:G) bases.
Next, the terminal sugar-phosphate is removed by an AP endonuclease (Ape1), leaving a 3’-OH terminal and an abnormal 5'-abasic terminus. The resulting gap is refilled by the 5’-deoxyribose-phosphodiesterase action of a DNA polymerase I (DNA polymerase beta in mammals), then the strands are re-ligated by DNA Ligase I or a complex of XRCC1 and LigIII.
An alternative BER pathway corrects errors involving more than one nucleotide. The Fen1 protein excises the long-patch structure that is produced by DNA polymerase strand displacement. This "long-patch" repair process is divided into two subpathways: a PCNA-stimulated, Pol-beta-directed pathway and a PCNA-dependent, Pol-delta/epsilon -directed pathway.
link to table - human DNA repair genes : diagram>BER
Biochemistry Overview : Molecular Genetics Overview : SITE MAP : HOME





































